Uploading data to DCOR
Prerequisites
DCKit: graphical toolkit for the management of RT-DC data (https://github.com/DC-analysis/DCKit/releases)
DCOR-Aid: GUI for managing data on DCOR (https://github.com/DCOR-dev/DCOR-Aid/releases)
Data preparation with DCKit
In many cases, you should not upload your experimental data right away to DCOR. There may be several reasons for that, such as missing metadata, uncompressed raw data, or log files that contain sensitive or unnecessary information (such as the user name of the person that recorded or processed the raw data). Please also note that DCOR only works with DC data in the HDF5 file format (.rtdc file extension).
DCKit to the rescue! In most cases, it is sufficient to to run your data through DCKit. Load the files in question, run the integrity check, complete or correct any missing or bad metadata keys and either convert the data to the .rtdc file format (for tdms data) or compress the data. You can verify that everything went as intended by running the integrity check for the newly generated files. If you are certain that you are not losing valuable information, you may also use the repack and strip logs option.

Fig. 3 DCKit user interface with one .rtdc file loaded that passed all integrity checks. DCKit can perform various tasks that are represented by the tool buttons on the right. Before uploading to DCOR, it is recommended to at least update the metadata such that the integrity checks pass.
Data upload with DCOR-Aid

Fig. 4 The DCOR-Aid setup wizard guides you through the initial setup.
To upload your data to a DCOR instance, you first need to create an account. When you start DCOR-Aid for the first time, you will be given several options.
If you select “Playground”, DCOR-Aid will create a testing account at https://dcor-dev.mpl.mpg.de for you. All data on that development is pruned weekly. You can use the DCOR-dev instance for testing.
If you select “DCOR”, you will have to manually register at dcor.mpl.mpg.de and generate an API key for DCOR-Aid in the DCOR web interface.
You can always run the setup wizard again via the File menu to e.g. switch from “playground” to the production DCOR server.
Once DCOR-Aid is connected to a DCOR instance, go to the Upload tab. The New manual upload tool button directs you to the metadata entry and resource selection process. It is also possible to upload pre-defined upload tasks (see next section).
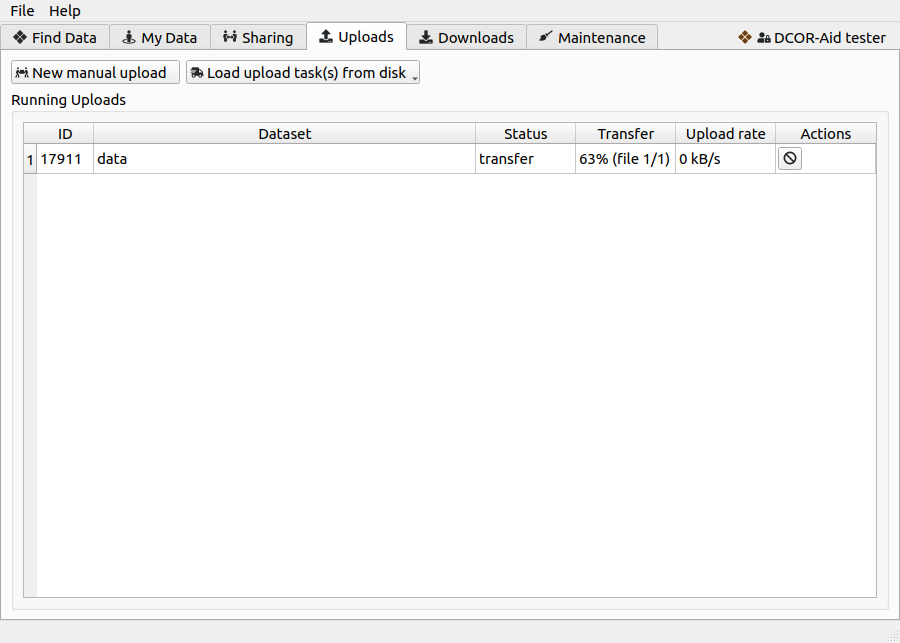
Fig. 5 The upload tab gives you the option to manually upload datasets or to load auto-generated DCOR-Aid upload task (.dcoraid-task) files via the buttons at the top. Queued and running uploads are then displayed in the table below.
Generating DCOR-Aid upload tasks
If you need a way to upload many datasets in an automated manner you can make use of .dcoraid-task files. These files are essentially upload recipes that can be loaded into DCOR-Aid in the Upload tab via the Load upload task(s) from disk tool button.
The following script upload_task_generation.py recursively
searches a directory tree for .rtdc files and generates .dcoraid-task files.
"""DCOR-Aid task creator
This script automatically generates *.dcoraid-task files recursively.
For each directory with *.rtdc files, an upload_job.dcoraid-task file
is generated. This task file can then be loaded into DCOR-Aid for the
actual upload. This script only serves as a template. Please go ahead
and edit it to your needs if necessary.
Changelog
---------
2021-10-26
- initial version
"""
import copy
import pathlib
import dclab
import dcoraid
#: Local directory to search recursively for .rtdc files
DATA_DIRECTORY = r"T:\Example_Data\Main_Directory"
#: List of file name suffixes of files to be included in the upload
#: (see :func:`dcoraid.CKANAPI.get_supported_resource_suffixes`)
DATA_FILE_SUFFIXES = [
"*.ini",
"*.csv",
"*.tsv",
"*.txt",
"*.pdf",
"*.jpg",
"*.png",
"*.so2",
"*.poly",
"*.sof",
]
#: Default values for the dataset upload
DATASET_TEMPLATE_DICT = {
# Should the datasets by private or publicly visible (optional)?
"private": False,
# Under which license would you like to publish your data (mandatory)?
"license_id": "CC0-1.0",
# To which DCOR circle should the dataset be uploaded (optional)?
"owner_org": "my-dcor-circle",
# Who is responsible for this dataset (mandatory)?
"authors": "Heinz Beinz Automated Upload",
}
#: Supplementary resource metadata
#: (see :func:`dcoraid.CKANAPI.get_supplementary_resource_schema`)
RSS_DICT = {
"cells": {
"organism": "human",
"cell type": "blood",
"fixed": False,
"live": True,
"frozen": False,
},
"experiment": {
"buffer osmolality": 284.0,
"buffer ph": 7.4,
}
}
def recursive_task_file_generation(path=DATA_DIRECTORY):
"""Recursively generate .dcoraid-task files in a directory tree
Skips directories that already contain a .dcoraid-task file
(This is important in case DCOR-Aid already imported that task
file and gave that task a DCOR dataset ID).
"""
# Iterate over all directories
for pp in pathlib.Path(path).rglob("*"):
if pp.is_dir():
generate_task_file(pp)
def generate_task_file(path):
"""Generate the upload_job.dcoraid-task file in directory `path`
A task file is only generated if the directory contains .rtdc
files.
"""
path = pathlib.Path(path)
assert path.is_dir()
path_task = path / "upload_job.dcoraid-task"
if path_task.exists():
print(f"Skipping creation of {path_task} (already exists)")
return
else:
print(f"Processing {path}", end="", flush=True)
# get all .rtdc files
resource_paths = sorted(path.glob("*.rtdc"))
# make sure they are ok
for pp in copy.copy(resource_paths):
try:
with dclab.IntegrityChecker(pp) as ic:
cues = ic.sanity_check()
if len(cues):
raise ValueError(f"Sanity Check failed for {pp}!")
except BaseException:
print(f"\n...Excluding corrupt resource {pp.name}",
end="", flush=True)
resource_paths.remove(pp)
# proceed with task generation
if resource_paths:
# DCOR dataset dictionary
dataset_dict = copy.deepcopy(DATASET_TEMPLATE_DICT)
# Set the directory name as the dataset title
dataset_dict["title"] = path.name
# append additional resources
for suffix in DATA_FILE_SUFFIXES:
resource_paths += path.glob(suffix)
# create resource dictionaries for all resources
resource_dicts = []
for pp in resource_paths:
rsd = {"path": pp,
"name": pp.name}
if pp.suffix == ".rtdc":
# only .rtdc data can have supplementary resource metadata
rsd["supplements"] = get_supplementary_resource_metadata(pp)
resource_dicts.append(rsd)
dcoraid.create_task(path=path_task,
dataset_dict=dataset_dict,
resource_dicts=resource_dicts)
print(" - Done!")
else:
print("\n...No usable RT-DC files!")
def get_supplementary_resource_metadata(path):
"""Return dictionary with supplementary resource metadata
You will probably want to modify this function to your liking.
"""
path = pathlib.Path(path)
assert path.suffix == ".rtdc"
supplements = copy.deepcopy(RSS_DICT)
# Here you may add additional information, e.g. if you want
# to add a pathology depending on the folder name of the
# containing folder:
#
# if path.parent.name.count("BH"):
# supplements["cells"]["pathology"] = "long covid"
#
return supplements
if __name__ == "__main__":
recursive_task_file_generation()